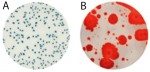
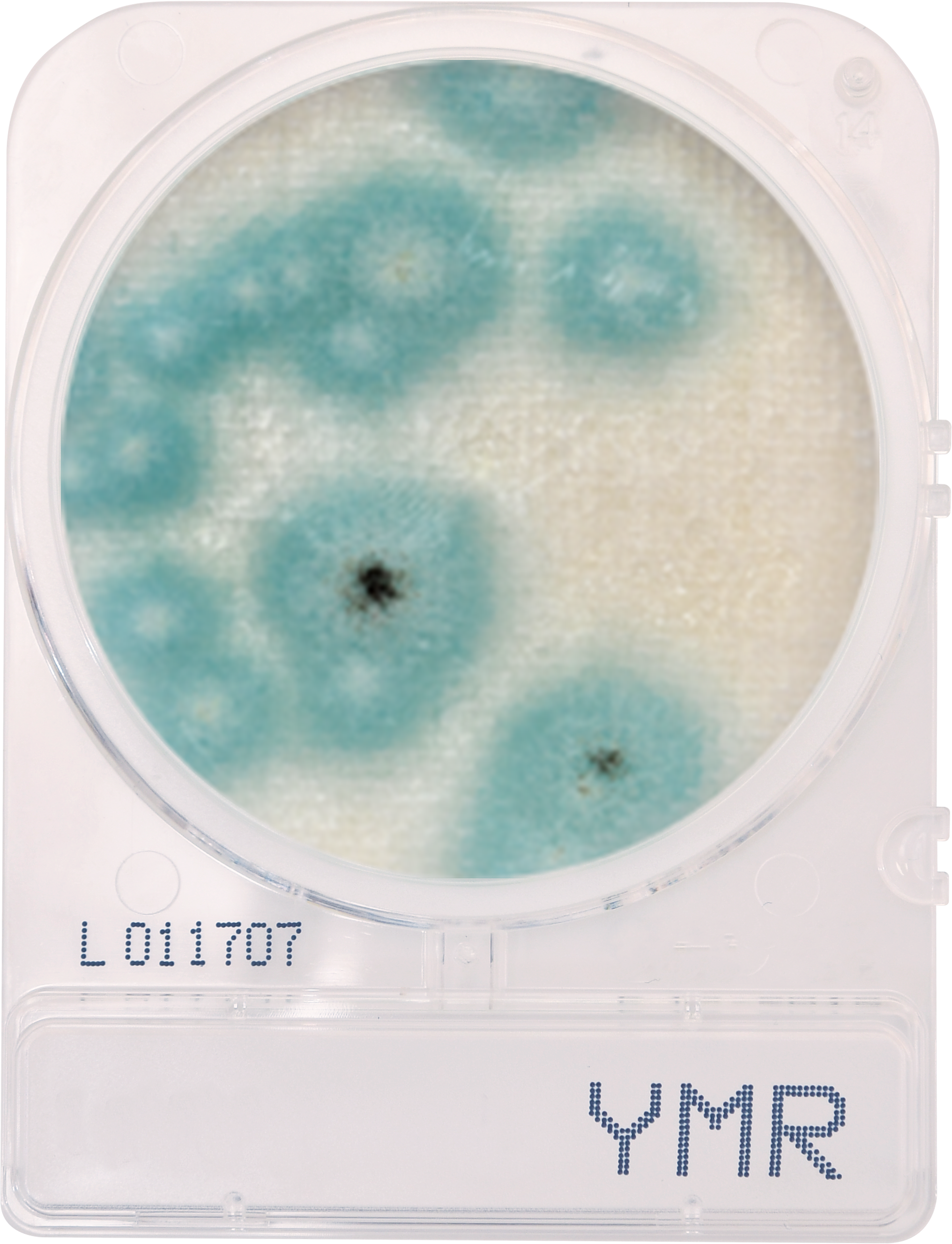
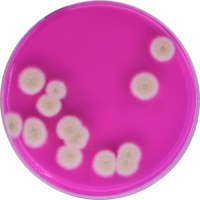
Cannabis testing to detect microbial contamination is complicated. It may not be rocket science, but it is life science, which means it’s a moving target, or at least, it should be, as we acquire more and more information about how the world we live in works. We are lucky to be able to carry out that examination in ever increasing detail. For instance, the science of genomics1 was born over 80 years ago, and just twenty years ago, genetics was still a black box. We’ve made tremendous progress since those early days, but we still have a long way to go, to be sure.
Much of that progress is due to our ability to build more accurate tools, a technological ladder, if you will, that raises our awareness, expertise, and knowledge to new levels. When a new process or technology appears, we compare it against accepted practice to create a new paradigm and make the necessary adjustments. But people have to be willing to change. In the cannabis industry, rapid change is a constant, first because that is the nature of a nascent industry, and second because in the absence of some universal and unimpeachable standard, it’s difficult to know who’s right. Especially when the old, reliable reference method (i.e. plating, which is basically growing microorganisms on the surface of a nutritional medium) is deeply flawed in its application to cannabis testing vs. molecular methods (i.e., quantitative polymerase chain reaction, or qPCR for short).

Plating systems have been used faithfully for close to 130 years in the food industry, and has performed reasonably well.2 But cannabis isn’t food and can’t be tested as if it were. In fact, plating methods have a host of major disadvantages that only show up when they’re used to detect cannabis pathogens. They are, in no particular order:
- A single plating system can’t enumerate a group of microorganisms and/or detect specific bacterial and fungal pathogens. This is further complicated by the fact that better than 98% of the microbes in the world do not form colonies.3 And there is no ONE UNIVERSAL bacterial or fungal SELECTIVE agar plate that will allow the growth of all bacteria or all fungal strains. For example, the 5 genus species of fungal strains implicated in powderly mildew DO NOT plate at all.
- Cannabinoids, which can represent 10-30% of a cannabis flower’s weight, have been shown to have antibacterial activity.4 Antibiotics inhibit the growth of bacteria and in some cases kill it altogether. Salmonella species & shiga toxin producing coli (STEC) bacteria, in particular, are very sensitive to antibiotics, which leads to either a false negative result or lower total counts on plates vs. qPCR methods.
- Plating methods cannot detect bacterial and fungal endophytes that live a part or all of their life cycle inside a cannabis plant.5,6 Examples of endophytes are the Aspergillus pathogens (A. flavus, A. fumigatus, A. niger, and A. terreus). Methods to break open the plant cells to access these endophytes to prepare them for plating methods also lyse these microbial cells, thereby killing endophytic cells in the process. That’s why these endophytes will never form colonies, which leads to either false negative results or lower total counts on plates vs. qPCR methods.
- Selective plating media for molds, such as Dichloran Rose-Bengal Chloramphenicol (DRBC) actually reduces mold growth—especially Aspergillus—by as much as 5-fold.This delivers false negative results for this dangerous human pathogen. In other words, although the DRBC medium is typically used to reduce bacteria; it comes at the cost of missing 5-fold more yeast and molds than Potato Dextrose Agar (PDA) + Chloramphenicol or molecular methods. These observations were derived from study results of the AOAC emergency response validation.7
- Finally, we’ve recently identified four bacterial species, which are human pathogens associated with cannabis that do not grow at the plating system incubation temperature typically used.8 They are Aeromonas hydrophila, Pantoea agglomerans, Yersinia enterocolitica, and Rahnella aquatilis. This lowers total counts on plates qPCR methods.
So why is plating still so popular? Better yet, why is it still the recommended method for many state regulators? Beats me. But I can hazard a couple of guesses.
First, research on cannabis has been restricted for the better part of the last 70 years, and it’s impossible to construct a body of scientific knowledge by keeping everyone in the dark. Ten years ago, as one of the first government-employed scientists to study cannabis, I was tapped to start the first cannabis testing lab at the New Jersey Dept. of Health and we had to build it from ground zero. Nobody knew anything about cannabis then.
Second, because of a shortage of cannabis-trained experts, members of many regulatory bodies come from the food industry—where they’ve used plating almost exclusively. So, when it comes time to draft cannabis microbial testing regulations, plating is the default method. After all, it worked for them before and they’re comfortable with recommending it for their state’s cannabis regulations.
Finally, there’s a certain amount of discomfort in not being right. Going into this completely new area—remember, the legal cannabis industry didn’t even exist 10 years ago—we human beings like to have a little certainty to fall back on. The trouble is, falling back on what we did before stifles badly needed progress. This is a case where, if you’re comfortable with your old methods and you’re sure of your results, you’re probably wrong.
So let’s accept the fact that we’re all in this uncharted territory together. We don’t yet know everything about cannabis we need to know, but we do know some things, and we already have some pretty good tools, based on real science, that happen to work really well. Let’s use them to help light our way.
References
- J. Weissenbach. The rise of genomics. Comptes Rendu Biologies, 339 (7-8), 231-239 (2016).
- R. Koch. 1882. Die Aetiologie der Tuberculose. Berliner Klinische Wochenschrift, 19, 221-230 (1882).
- W. Wade. Unculturable bacteria—the uncharacterized organisms that cause oral infections. Journal of the Royal Society of Medicine, 95(2), 91-93 (2002).
- J.A. Karas, L.J.M. Wong, O.K.A. Paulin, A. C. Mazeh, M.H. Hussein, J. Li, and T. Vekov. Antibiotics, 9(7), 406 (2020).
- M. Taghinasab and S. Jabaji, Cannabis microbiome and the role of endophytes in modulating the production of secondary metabolites: an overview. Microorganisms 2020, 8, 355, 1-16 (2020).
- P. Kusari, S. Kusari, M. Spiteller and O. Kayser, Endophytic fungi harbored in Cannabis sativa L.: diversity and potential as biocontrol agents against host plant-specific phytopathogens. Fungal Diversity 60, 137–151 (2013).
- K. McKernan, Y. Helbert, L. Kane, N. Houde, L. Zhang, S. McLaughlin, Whole genome sequencing of colonies derived from cannabis flowers & the impact of media selection on benchmarking total yeast & mold detection tools, https://f1000research.com/articles/10-624 (2021).
- K. McKernan, Y. Helbert, L. Kane, L. Zhang, N. Houde, A. Bennett, J. Silva, H. Ebling, and S. McLaughlin, Pathogenic Enterobacteriaceae require multiple culture temperatures for detection in Cannabis sativa L. OSF Preprints, https://osf.io/j3msk/, (2022)