Hop latent viroid (HLVd) has gained attention as the molecular cause of “dudding disease” and is causing significant economic losses in the cannabis industry.1,2 Estimates indicate that upwards of 4 billion dollars of market value are lost each year to this pathogen alone.3 The impact of HLVd on cannabis plants necessitates the development and implementation of effective pathogen diagnostics to mitigate its spread and minimize crop damage. With collaborative research efforts, we can gain valuable insights into the characteristics, spread, symptoms and preventive measures associated with HLVd in the cannabis industry.
Viroids: A Brief Overview

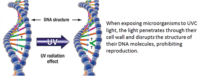
Viroids are unique infectious agents composed solely of genetic material, distinct from viruses. Unlike viruses, viroids lack a protective protein layer and solely rely on the host plant for replication and spread. Their stability and ability to persist in various environments make viroids a formidable threat to plant health.
Hop Latent Viroid: Origin and Global Spread
Hop latent viroid was initially identified in hop plants in 19884 and was found to be largely asymptomatic in this crop. Consequently, HLVd has spread worldwide, mostly unchecked by the hops industry. This pathogen has been identified on most continents and in some fields more than 90% of hops plants are infected.5 Hop latent viroid very likely jumped from hops into cannabis, due to similar genetics. The timing and mechanism of cross-species transmission to cannabis remains unknown, but the prevalence of HLVd suggests this viroid has been circulating within cannabis for an extended period. Data collected at TUMI Genomics indicates that HLVd is present in all states in the United States where cannabis is legal as well internationally including; Canada, the United Kingdom, France, the Netherlands, Thailand, Austria and Switzerland.
Symptoms and Impacts on Cannabis Plants

HLVd exhibits a wide range of symptoms, which can vary from severe to subtle, affecting the growth, leaf development, flower quality and overall vitality of cannabis plants. Understanding these symptoms is crucial for timely diagnosis and appropriate disease management strategies. However, HLVd can also present asymptomatically, especially in vegetative plants. The only way to determine if your plants are infected is by routine molecular testing.
Modes of Transmission
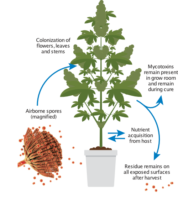
Mechanical Transmission: HLVd primarily spreads mechanically through contact with infected sap during activities like trimming and handling. Additionally, transmission through contaminated water and the potential role of insects, fungal pathogens and seeds in spreading HLVd have also been observed.
Seed Transmission: Although no published studies exist in cannabis describing the frequency of seed transmission, HLVd does transmit through seeds in hop plants at a rate of around 8%.7 Preliminary studies performed by TUMI Genomics in collaboration with EZ-genetics suggest cannabis seed transmission does occur at variable rates depending on strain and level of infection of the parent plants.
Water Transmission: It has also been observed that viroids are in high concentration in the roots8 and can move from the root into runoff water.9 Plants sharing a common water source with infected plants, such as recirculating water systems or flood and drain procedures, are at risk for transmission of the viroid.
Insect and Other Vector Transmission: The jury is still out as to whether or not insects can transmit HLVd. However, multiple viroids are transmitted via insects, so it is likely that HLVd insect transmission occurs. Recent studies also indicate that fungal pathogens, like Fusarium, can transmit viroid infections.6 While pathogenic fungus is a major concern for cannabis growers in its own right, limiting the prevalence and spread of fungal pathogens in your facility could help limit hop latent viroid transmission as well.
Therefore, implementing proper sanitation practices and limiting pest access can help minimize transmission risks.
Preventive Measures
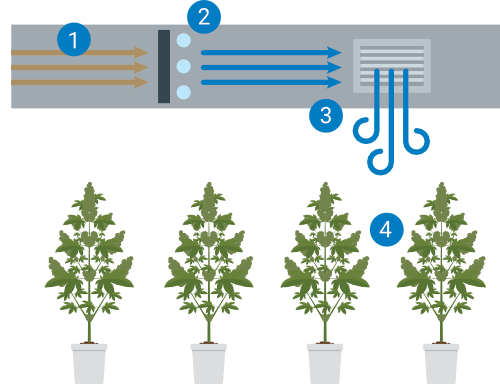
Prevention plays a vital role in safeguarding cannabis crops against HLVd. The STOP program, developed by TUMI Genomics, offers a comprehensive approach that includes maintaining a Sterile environment, Testing mother plants regularly, Organizing the facility to minimize pathogen spread, and Protecting the facility’s borders from introduction of infected plant material, insects and contaminated water. More details on these preventative measures can be found here.
Pathogen Diagnostics
Protecting your plants from hop latent viroid requires accurate identification and removal of infected plants before the infection spreads to other plants. To accomplish this, several critical factors should be considered:
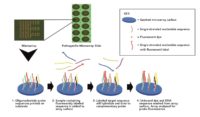
Type of test: HLVd and all viroids can only be detected by a molecular test (a test that detects the presence of DNA/RNA). Among common molecular tests, PCR is generally the most sensitive and accurate method. PCR can provide both a diagnosis and an approximate viroid level, allowing informed management decisions. Other types of molecular tests, such as LAMP and RPA, can formally be as sensitive as PCR, but the classic versions of these assays often suffer from false positive/negative results, reducing accuracy.
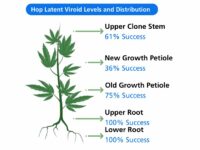
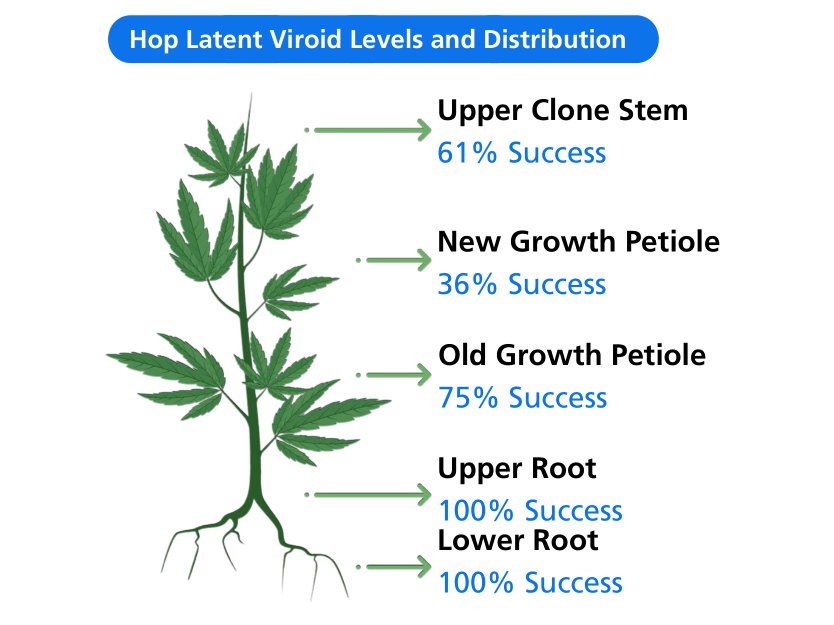
Tissue type: An important consideration for HLVd detection is the plant tissue selected for testing, especially when identifying low-level or early infections when HLVd is not yet systemic. Studies completed by TUMI Genomics and others show root tissue contains the highest levels of HLVd and is the most reliable tissue for detection of viroid infection. While upper root tissue appears to contain the highest levels of viroid, roots from anywhere in the root ball are predictive of infection. Samples taken from the leaves/foliage tend to have lower levels of viroid and may produce false negative results.
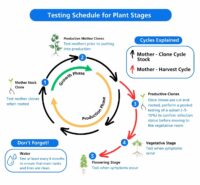
Testing frequency: Routine pathogen testing is standard practice in general agriculture and is critical to maintain a healthy cannabis crop. Testing of mother plants every 4-6 weeks for economically critical pathogens (such as HLVd) will help ensure a successful run and a high-quality product.
Disinfection Methods
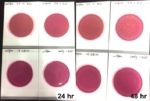
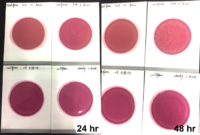
Studies have shown that viroids can remain infectious for longer than 24 hours on most common surfaces11 and 7 weeks in water.10 Making effective disinfection methods essential to limit the spread of HLVd. While common disinfectants like alcohol and hydrogen peroxide are ineffective against viroids, a 10% bleach solution has shown efficacy in destroying HLVd. Proper tool sterilization practices, such as soaking tools in bleach for 60 seconds, are crucial to prevent transmission during plant handling.

Hop latent viroid poses a significant threat to the cannabis industry, leading to substantial economic losses. Timely and accurate pathogen diagnostics, along with stringent preventive measures, are essential for minimizing the impact of HLVd. Regular testing, proper disinfection protocols and adherence to pathogen prevention programs can help ensure the health and vitality of cannabis crops in the face of this global pandemic.
References
- Bektas, A., et al. “Occurrence of Hop Latent Viroid in Cannabis Sativa with Symptoms of Cannabis Stunting Disease in California.” APS Journals, 21 Aug. 2019, doi.org/10.1094/PDIS-03-19-0459-PDN.
- Warren, J.G., et al. “Occurrence of Hop Latent Viroid Causing Disease in Cannabis Sativa in California.” APS Journals, 21 Aug. 2019, doi.org/10.1094/PDIS-03-19-0530-PDN.
- Cooper, Benjie. “Hop Latent Viroid Causes $4 Billion Cannabis Industry Loss – Candid Chronicle.” Candid Chronicle – Truthful, Straightforward, Blunt Cannabis News, 16 Aug. 2021, candidchronicle.com/hop-latent-viroid-causes-4-billion-cannabis-industry-loss/.
- Puchta H, Ramm K, Sänger HL. The molecular structure of hop latent viroid (HLV), a new viroid occurring worldwide in hops. Nucleic Acids Res. 1988 May 25;16(10):4197-216. doi: 10.1093/nar/16.10.4197. PMID: 2454454; PMCID: PMC336624.
- Faggioli, Franceso, et al. “Geographical Distribution of Viroids in Europe.” Viroids and Satellites, 31 July 2017, www.sciencedirect.com/science/article/abs/pii/B9780128014981000449#bib47.
- Wei S, Bian R, Andika IB, Niu E, Liu Q, Kondo H, Yang L, Zhou H, Pang T, Lian Z, Liu X, Wu Y, Sun L. Symptomatic plant viroid infections in phytopathogenic fungi. Proc Natl Acad Sci U S A. 2019 Jun 25;116(26):13042-13050. doi: 10.1073/pnas.1900762116. Epub 2019 Jun 10. PMID: 31182602; PMCID: PMC6600922.
- Singh RP. The discovery and eradication of potato spindle tuber viroid in Canada. Virus disease. 2014 Dec;25(4):415-24. doi: 10.1007/s13337-014-0225-9. Epub 2014 Dec 2. PMID: 25674616; PMCID: PMC4262315.
- Jama, Aisha, et al. TUMI Genomics, Fort Collins, CO, 2022, Hop Latent Viroid Levels and Distribution in Cannabis Plant Tissue.
- Mackie AE, Coutts BA, Barbetti MJ, Rodoni BC, McKirdy SJ, Jones RAC. Potato spindle tuber viroid: Stability on Common Surfaces and Inactivation With Disinfectants. Plant Dis. 2015 Jun;99(6):770-775. doi: 10.1094/PDIS-09-14-0929-RE. Epub 2015 May 15. PMID: 30699527.
- Mackie AE, Coutts BA, Barbetti MJ, Rodoni BC, McKirdy SJ, Jones RAC. Potato spindle tuber viroid: Stability on Common Surfaces and Inactivation With Disinfectants. Plant Dis. 2015 Jun;99(6):770-775. doi: 10.1094/PDIS-09-14-0929-RE. Epub 2015 May 15. PMID: 30699527.
- Mackie AE, Coutts BA, Barbetti MJ, Rodoni BC, McKirdy SJ, Jones RAC. Potato spindle tuber viroid: Stability on Common Surfaces and Inactivation With Disinfectants. Plant Dis. 2015 Jun;99(6):770-775. doi: 10.1094/PDIS-09-14-0929-RE. Epub 2015 May 15. PMID: 30699527.